Cancer Measurement Technology and Data Science
The Cancer Measurement Technology and Data Science (CMD) program inspires fundamental advances in measurement technologies and data science, harnessing them to accelerate cancer research and provide clinical information faster, earlier, and when needed. The CMD program’s activities involve fundamental investigations into measurement and data sciences, engineering of novel technologies (i.e., imaging and molecular measurement devices) and data tools (i.e., big data, simulation, and interventional tools), the advancement of integrated systems for point of care deployment, and development of protocols, policies, and simulation tools. The program also emphasizes the introduction of concepts from other diseases to benefit cancer-related research and applications, as well as translation of cancer-inspired technologies to other diseases and applications.
Imaging
Bringing imaging diagnostics, especially rapid and molecularly-inspired, to the patient.
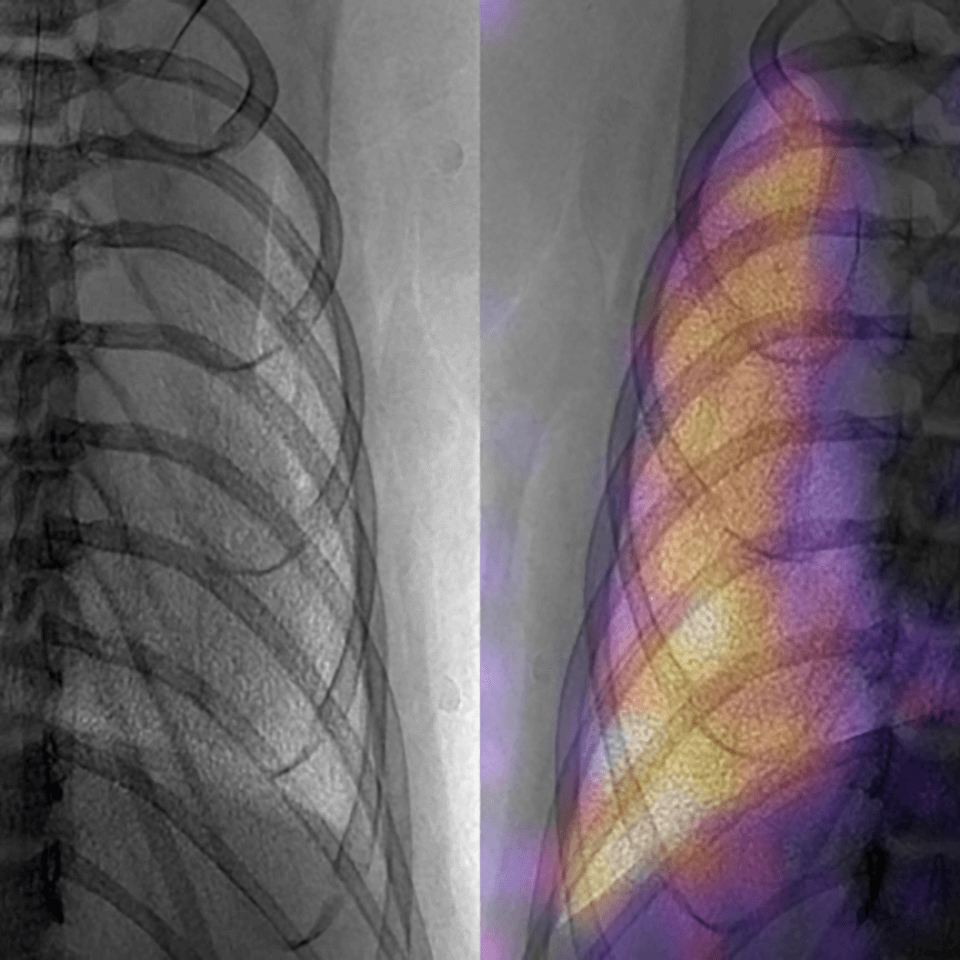
Traditionally, diagnostic imaging is conducted in a specialized laboratory. Here, the goal is to bring diagnostic imaging closer to the patient and add information that goes beyond traditional capabilities via innovative molecular contrast, increased information content, and the use of computing. The activities to advance this theme will:
- Inspire fundamental advances in theory and computational methods to enhance imaging, including higher performance of traditional clinical tools and development of new capabilities.
- Drive optical imaging technology to greater information content, faster imaging, and clinical use.
- Enable real-time, intraoperative imaging systems.
- Enable new technologies for pathology, including the development of stain-free, slide-free molecular digital histopathology and technologies for imaging the microenvironment.
- Generate the “imaging connectome” – foundational sets of transformations across imaging modalities, imaging physics, and imaging scales.
Molecular Measurement
Realizing molecular measurement technologies for early detection, treatment monitoring, and personalized therapy.
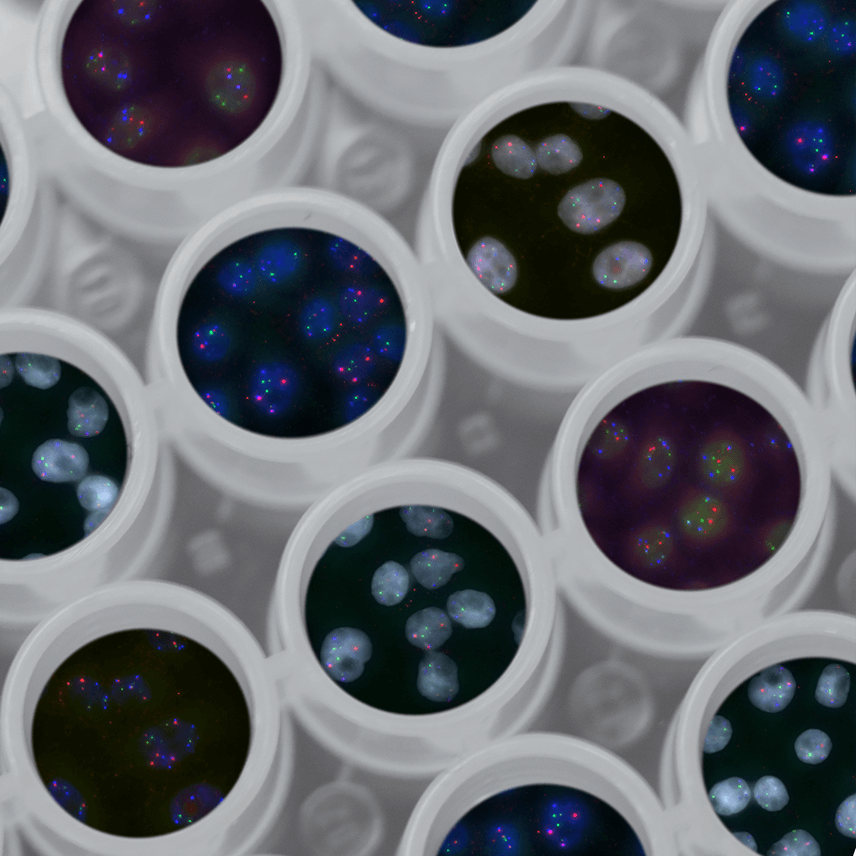
Increasingly, cancer molecular analyses are revealing new biomarkers that can illuminate disease-driving mutations. Similarly, clinical studies are showing how specific biomarkers can be used to quantify the effects of chemotherapy or surgery on a tumor. Making fundamental scientific progress towards ultra-sensitive, ultra-selective, and multiplexed biosensor technologies, we envision minimally invasive, simple, frequent, and inexpensive monitoring of the cancer transcriptome. Our approach to liquid biopsy will enable routine surveillance for early detection, tailoring therapy selection to the genomic characteristics of an individual’s disease and monitoring for recurrence. The goal of this theme is to utilize genomic biomarker testing in support of managing cancer through a combination of early detection, personally tailored treatment, and rapid therapy modification to shifting genomic characteristics via:
- Fundamental progress in enhanced sensing mechanisms with single molecule precision and ultra-high selectivity. We will also emphasize the development of microfluidic devices for automated extraction and pre-concentration of nucleic acids, exosomes, and protein targets from complex sample media (whole blood, urine, saliva, stool) as standalone technologies and adjuncts to measurement technologies.
- Development of assay methods and instrumentation to enable analysis that can be performed in point of care settings.
- Tools for monitoring the pre-cancer human transcriptome to establish personalized baselines that also inform upon the genomic effects of nutrition, behavior (exercise), and the environment.
- Association of cancer genomic response with biomarkers that measure co-morbidities (heart disease, infectious disease, liver disease) and the microbiome.
- Bringing technologies for measurements to the patient. These will include rapid point-of-care personalized detection of cancer and point-of-administration sensing of drug efficacy and wearable sensors for continuous cancer surveillance and treatment response monitoring.
Computational Biology and Engineering
Democratizing molecular analysis, diagnosis, and decision making by enabling state of the art computing and artificial intelligence.
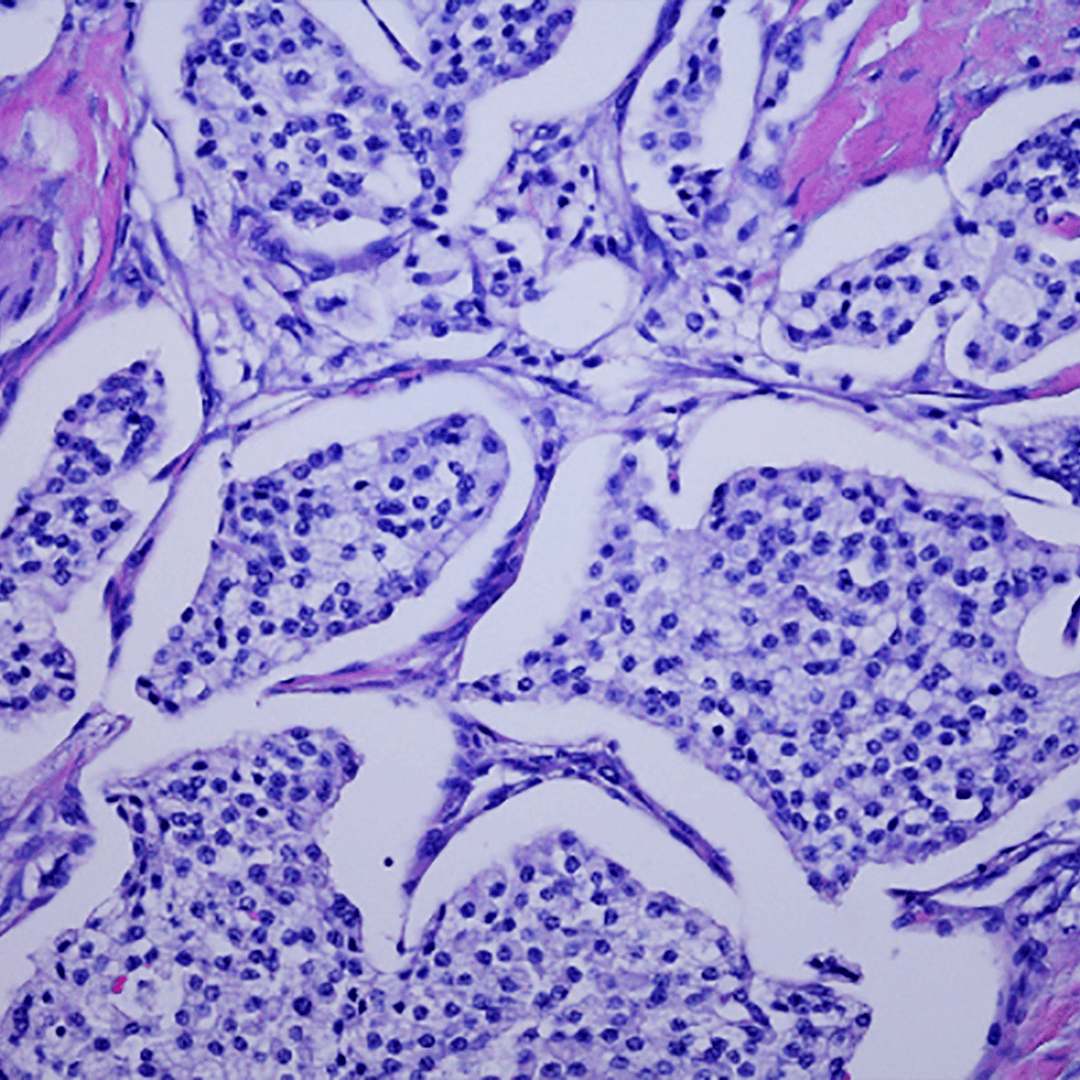
Computational approaches and methods undergird a significant portion of research within the CCIL. We will focus on advances in computational methods, hardware to support these efforts and the development of software for wide use in basic research and cancer applications. Our computational methods and technologies can broadly improve capabilities in modeling, measurement, and analysis. In particular, hardware advances are augmented and enhanced when computation, new performance, reliability, consistency, and accuracy are realized. Our approach is to co-develop hardware and computing systems that will revolutionize not only the availability of data, but also our capabilities to transform data to knowledge, and knowledge to diagnosis and decision making. We will also use our expertise in computing to drive collaborations that would otherwise not be possible. Activities in this theme include:
- Fundamental advances in computing, statistics, and applied mathematics that can be applied to cancer.
- Development of machine learning and artificial intelligence algorithms to provide actionable information through analysis of biomarkers, complemented by data from medical imaging, population statistics, patient genome, environmental factors, and patient medical record. Predictive AI for personalized cancer therapies and guided interventions (e.g. in robotic surgeries) will also be emphasized.
- Development of big data to knowledge software and tools, including biomarker discovery and rapid assessment of molecular targets for drug discovery.
- Computational reconstruction methods, from imaging instruments to reconstructing biological structures.
- Novel methods to store, handle and curate data.
Cancer Discovery Platforms Bridging the Engineering-Biology Continuum Program
The guiding principle of the Cancer Discovery Platforms Bridging the Engineering-Biology Continuum (CDP) Program is to develop platforms to facilitate the study of pathophysiological processes linked to cancer progression, drug efficacy, and malignant cellular vulnerabilities through the use of sophisticated model systems. We aim to develop and characterize state-of-the-art platforms that model the chaos and contextual complexity of the cancer microenvironment to accelerate the identification, targeting, and validation of new anticancer therapies. Fundamental scientific inquiry and early-stage development of molecules, models, and methods form the foundation of our efforts in drug discovery and translation of these discoveries to industry, expediting their application to patients.
Pathways and Mechanisms
Enabling biological insight, focused on cancer pathways and mechanisms tied to outcomes.
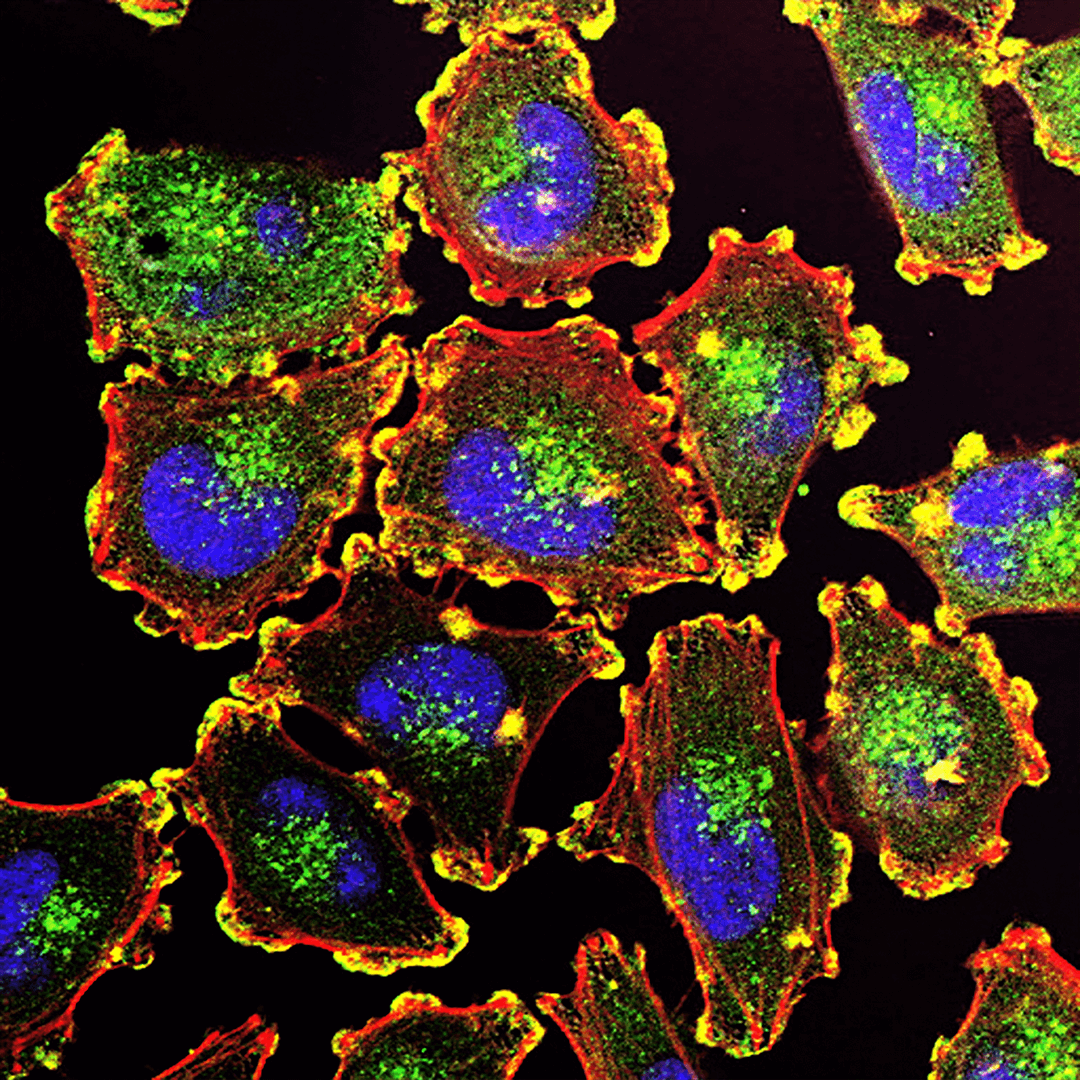
Researchers
- Sayeepriyadarshini Anakk
- Milan Bagchi
- Jie Chen
- Catherine Christian-Hinman
- Matthew Dean
- John Erdman
- Jodi Flaws
- Brian Freeman
- Rex Gaskins
- William Helferich
- Raven Huang
- Keith Jarosinski
- Hong Jin
- Auinash Kalsotra
- Benita Katzenellenbogen
- Brett Loman
- Zeynep Madak Erdogan
- Ruby Mendenhall
- Satish Nair
- Erik Nelson
- Kannanganattu Prasanth
- Supriya Prasanth
- Huanyu Qiao
- Jason Ridlon
- Andrew Steelman
- Patrick Sweeney
- Kevin Van Bortle
- Bo Wang
- Nicholas Wu
It is essential to leverage the strengths of the CCIL to elucidate targetable pathways linked to cancer pathophysiology. Our ongoing efforts will improve mechanistic insight regarding genetic mutations, RNA regulation, as well as receptor activation and signal transduction. There are significant opportunities to leverage approaches and platforms to investigate reciprocal relationships within the tumor microenvironment related to metabolism, immune targeting, and cellular crosstalk. We are focused on advancing:
- Fundamental studies on RNA biology that relate to cancer, including lncRNA.
- Metabolic dependencies, restrictions, and adaptations.
- Connecting nutrition, lifestyle, and microbiome to cancer biology and progression.
- Cancer-immune axis and immunotherapy.
- Identifying dynamics of reparative and malignant remodeling within the tumor microenvironment.
Drug Discovery
Accelerating therapeutic discovery by advancing synthesis, computational discovery, and high-throughput screening.
The University of Illinois has long-standing expertise in organic chemistry, including natural products chemistry. Increasingly sophisticated efforts to identify targetable pathways requires innovations regarding the efficient development of next generation precision therapies against cancer, including small molecules, immunotherapies, and designer nanoparticles. We envision the integration of high-throughput screening modalities with computationally-aided acceleration of drug design to identify novel single and combination therapies against cancer.
Major directions include:
- Fundamental research into organic synthesis relevant to cancer, including that of natural products.
- Discovery of new compounds by high-throughput screening and optimization.
- Systems engineering and computational acceleration of drug discovery by modeling.
- Target development through genomic and proteomic interrogation of cancer cells and tissues.
- Innovate drug delivery strategies through engineering technologies, including nanotechnology approaches.
Natural and Model Systems
Developing model systems for cancer research through engineered, naturally-occurring, and inducible constructs.
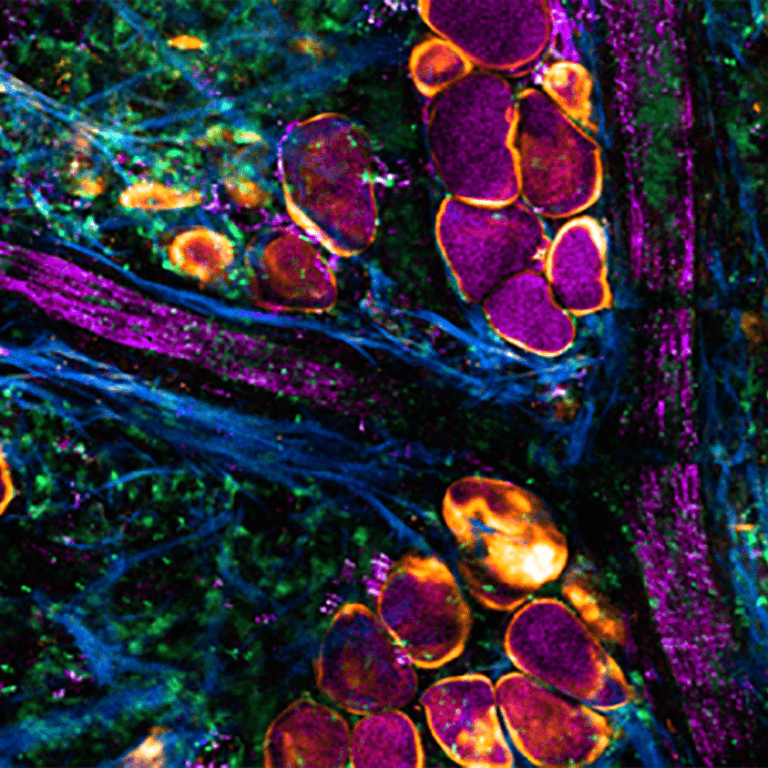
Novel therapeutic approaches are traditionally studies in two-dimensional culture or using inbred mouse models. Our approach is to validate an entirely new development pipeline to accelerate the evaluation and validation of new therapeutic interventions. We are developing multidimensional platforms that replicate cellular and microenvironmental complexity of the tumor to identify patterns of progression, invasion and metastasis, and drug resistance essential for evaluating new therapeutic drugs. Spontaneously arising and inducible veterinary oncology large animal models and technologies to leverage primary patient specimens will accelerate the evaluation of novel drug compounds.
Major directions include:
- Use of large animal models (e.g., companion animals and OncoPig) to accelerate drug development).
- Develop and translate tumor models that facilitate spatial and temporal analysis of patient biospecimens via quantitative molecular diagnostics and molecular sensing platform technologies.
- Fundamental progress in engineered platforms, especially those that replicate vascular remodeling and angiocrine signaling in the tumor microenvironment.
- Biofabrication innovations pertaining to 3D printing, microfluidic patterning, and the use of adaptable biomaterial chemistries to replicate dynamic tissue environments.
