Brian Cunningham, left, is joined by research team members Priyash Barya, Skye Shepherd, and Yanyu Xiong
Researchers from the Cancer Center at Illinois (CCIL) Program Leader Brian Cunningham’s lab in collaboration with researchers at Washington University have demonstrated a new capability to detect and count individual biomolecules at low concentrations. This technology may significantly improve the efficacy of current cancer detection and measurement methods.
Biomarkers play a significant role in diagnostics because their presence and quantity correlate to the presence of diseases like cancer. Low signal-to-noise ratios hamper current biomarker detection technology, posing a threat to cancer diagnostics, especially for patients whose cancer therapies can cause a reduction of detectable biomarker quantities in their system.
“To address this limitation, our team designed and developed a technique that leverages photonics to amplify the signal and enhance the testing capabilities. Our goal was to provide a more sensitive protein detection platform, ultimately leading to better diagnosis and treatment of diseases,” said project member Priyash Barya.
Cunningham’s team successfully met their goal, achieving an unprecedented ultrasensitive detection limit for protein biomarkers as low as 100 femtograms (fg) per milliliter (ml) (a femtogram is 10-15 grams).
“This kind of detection limit is not achievable with other technologies, except for one other that uses a complex procedure and expensive instrumentation that isn’t viable for point-of-care cancer diagnostics,” said Cunningham. “The ability to go down to concentrations this low is like pulling back the waters on the beach. Formerly hidden things are now visible. This method provides us with more information for early cancer detection and also for seeing the effects of cancer therapy on protein biomarkers that otherwise wouldn’t be known.”
How does the new method work? “It’s like chocolate and peanut butter,” said Cunningham. “Our new biomarker detection technology combines two great things to make something even better.”
First, the team at Washington University led by Prof. Srikanth Singamaneni created plasmonic fluors (PF), or gold nanoparticles decorated with fluorescent dye molecules. These PFs absorb light efficiently from a specific laser wavelength that couples the energy into the fluorescent dye molecule. The PF then becomes a tag for protein molecules. While PFs are ordinarily bright, researchers still need an expensive microscope to see them. But Cunningham’s team wanted to detect individual PFs, and to accomplish this feat, they performed the detection on the top of a photonic crystal (PC) designed to capture the same wavelength of light (in the laser) also used to excite the PF. The PF then piggybacks onto the PC with the biomarker present. Then the team efficiently couples more energy into the PF, increasing its brightness and enabling single-molecule detection.
A second novel effect gave the researchers an advantage – the emitted photons followed a well-defined path and direction dictated by the PC. The team chose a microscope with an appropriately wide aperture to efficiently capture this light. When combining this ability to visualize the brightness with the ability to see individual biomolecules, you get that ‘chocolate and peanut butter’ dynamic duo that distinguishes this new biomarker detection technology (see figure below).
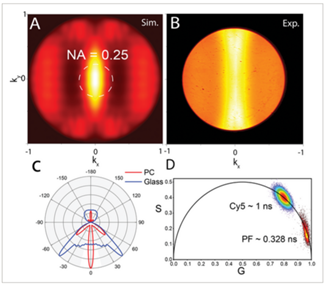
Figure 1: Light captured efficiently using the Cunningham lab’s novel ultrasensitive protein detection method appearing in the center of the image, like the “eye of Sauron.”
While it sounds complex, this new technology is inexpensive to manufacture, and the detection methodology is quite simple, requiring no additional enzymes for amplification, such as those used with PCR tests.
The Cunningham lab also recently published research on a new biomarker detection method for mRNA molecules, another type of biomarker that provides different information about cancer. With the mRNA biomarker detection, an amplification effect allowed even lower detection limits. One might wonder if these new biomarker detection methods work together. “We are thinking about that, actually. For example, we could combine light-emitting nanoparticles with these amplification methods,” said Cunningham.
“When we began this research endeavor two years ago, I was very interested in the concept of plasmonic and photonic coupling and its role in fluorescence enhancement,” said Priyash. “However, what particularly motivated me was the prospect of utilizing our method to enhance the sensitivity of current diagnostic capabilities. The possibility of contributing to advancements in diagnostics and potentially improving patient outcomes was a driving force for my involvement in this research.”
Where do things go from here for the team’s research?
“Our method could be easily translated to a straightforward and affordable optical instrument that promises to improve the sensitivity of present diagnostic capabilities. With enhanced sensitivity and accuracy in protein detection, it has the potential to contribute to early disease diagnosis and enable more efficient and timely treatments,” said Priyash.
Now that the team has demonstrated the principle of this new method, they will seek clinical collaborators who will pilot this technology’s capability for biomarker detection at small concentrations. The team will also apply for additional funding and continue collaborations with Washington University in St. Louis, MO.
Co-first authors Priyash Barya, Skye Shepherd, and Yanyu Xiong and principal investigator Brian Cunningham published these findings in the paper “Photonic-Plasmonic Coupling Enhanced Fluorescence Enabling Digital-Resolution Ultrasensitive Protein Detection” in Small, a leading multidisciplinary journal covering a broad spectrum of topics at the nano- and microscale.
Editor’s notes:
Brian Cunningham is a Cancer Center at Illinois (CCIL) program leader for the Cancer Measurement Technology and Data Science program, the Intel Alumni Endowed Chair of the Department of Electrical and Computer Engineering, a professor of bioengineering, and the Holonyak Micro and Nanotechnology Lab. Cunningham is also the Director of the Center for Genomic Diagnostics theme at the Carl R. Woese Institute for Genomic Biology.
To contact Brian Cunningham, email bcunning@illinois.edu
The work was supported by the National Science Foundation (1900277), the National Institutes of Health (NIH R01 5R01CA227699-03), and the Cancer Center at Illinois (CCIL).
The paper “Photonic-Plasmonic Coupling Enhanced Fluorescence Enabling Digital-Resolution Ultrasensitive Protein Detection” is available online. doi.org/10.1002/smll.202207239
Written by Jonathan King, CCIL Communications Team

